Script(s) of the Day: Exporting & importing binary masks
Export annotated regions as binary masks. Also, create new annotations from binary masks.
Inevitably, I got off-track with Script of the Day, and missed it yesterday.
So here are two scripts today, handling related tasks:
- Exporting annotations as binary masks
- Importing the binary masks again, creating new QuPath annotations
Problem
QuPath can be handy simply as a whole slide image viewer with annotation capabilities - especially for people who want to take care of the analysis elsewhere themselves.
But it helps to be able to get annotations into and out of QuPath.
One way is to work with the vertices of the annotations (e.g. rectangles, polygons). That approach will probably be the subject of a later post.
Another way is to create binary masks for annotations, which can then be reused elsewhere.
Because a particularly interesting use-case for this is in the generation of training data to create new algorithms, it helps to also be able to export pixel data from the whole slide image corresponding to the masks, at the same resolution.
Solution
The first script below shows how to start from one or more QuPath annotations…
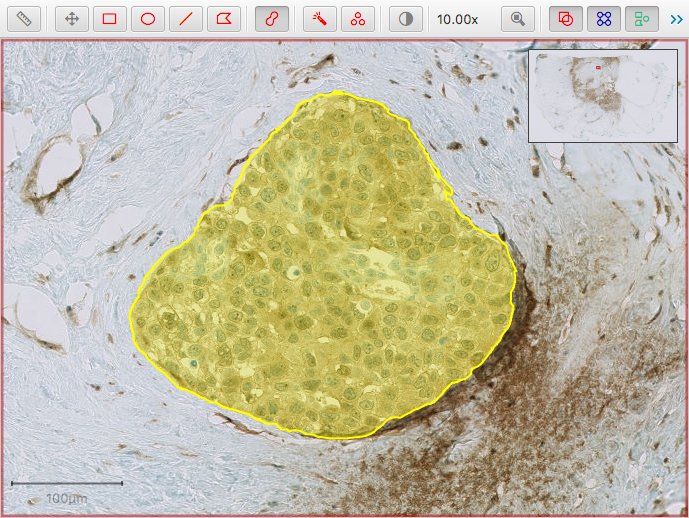
…and export these as binary masks…
-mask.png)
…optionally alongside the corresponding region of the image.
.jpg)
The script also encodes the following in the file name of the binary mask:
- the original image name
- the classification of the annotation
- the downsample value dictating the export resolution
- the bounding box of the region that was exported
This supports the job of the second script: to generate QuPath annotations based on binary images that use similar naming conventions.
Of course, this means that regions generated by image processing using any software capable of writing binary images can be imported into QuPath for visualization in the context of the whole slide image. It’s just a matter of adapting the script below, or using the same file naming convention.
Together, these scripts give two ways in which QuPath can help support researchers developing novel deep learning algorithms for pathology:
- Generating training data
- Visualizing results on whole slide images
However, there are a lot more ways in which QuPath can be useful, beyond these scripts.
Hopefully, over time this blog can describe in more detail how QuPath can be a useful algorithm development platform, even for people who ultimately might not want to run their algorithms through QuPath.
Entire scripts
Neither of the scripts below has been very extensively tested - so it’s worth looking through the code to check if they meet your needs, or require some adaption. There are also a few variables that could be adjusted to customize the behavior easily.